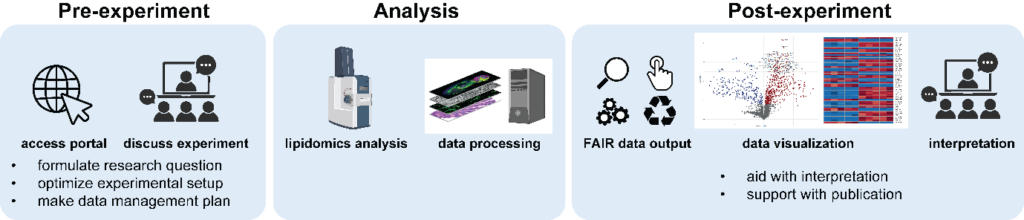
Pre-experiment, NExTLi screens projects to align research questions, experimental setups, and samples, crucial as many users lack lipidomics expertise but also will provide guidance to data management. This process prevents unnecessary costs and facility overuse from likely unsuccessful projects. Post-experiment, users receive standard data reports and custom visualizations / data representations, but often need assistance in data interpretation, contextualizing it with lipid metabolism and their research questions. Facility experts will help in writing papers, ensuring data quality, benefiting users to get most of the data!
NExTLi operates in seamless alignment with the Core Facility Metabolomics (CFM)) within the Laboratory Genetic Metabolic diseases at the Amsterdam UMC. As we work on developing a specialized NExTLi portal complete with detailed instructions and resources, we kindly request that you reach out through the established channels of the CFM for any inquiries or support needs. In the table below we list the current services.
NExTLi services
| Service | Specification of the service |
| 2D-lipidomics | Orbitrap-based lipidomics: UPLC-separation, exact m/z determination, data processing and reporting. |
| 3D/4D-lipidomics | timsTOF Pro 2-based or Orbitrap-based (Exploris 240 only, using AcquireX) lipidomics: UPLC-separation, with/without ion mobility (CCS), exact m/z determination and fragmentation for ultimate lipidomics coverage/annotation certainty, data processing and reporting |
| 2D/3D/4D-fluxomics | Orbitrap-based or timsTOF Pro 2-based lipidomics using stable isotope labeled precursors to follow lipid metabolism, custom data processing and reporting |
