NExTLi
The National Expert Center for Translational Lipidomics
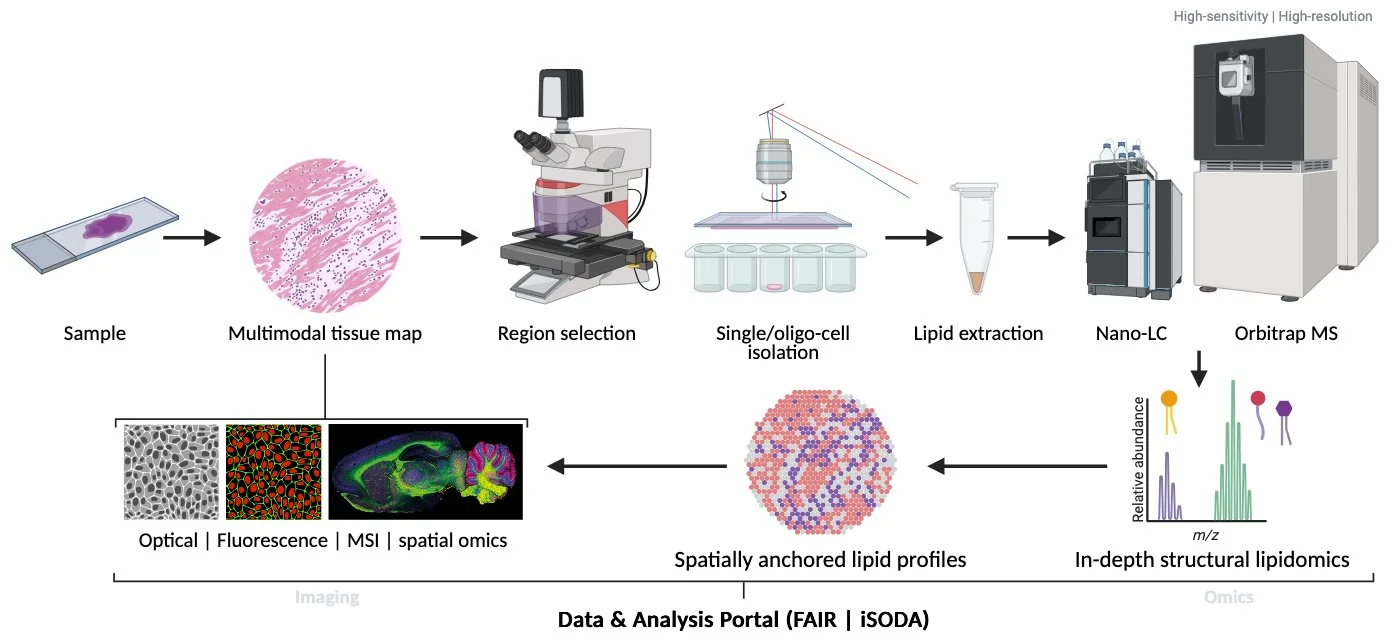
A coordinated Dutch research infrastructure for mechanistic lipid biology — integrating high-resolution structural lipidomics, spatially informed sampling, and tracer-based metabolic flux analysis within a single, expert-guided workflow. NExTLi enables lipid composition, organization, and metabolism to be studied with molecular specificity, spatial context, and dynamic resolution, supported by FAIR-by-design data delivery. The figure below illustrates how a tissue sample moves through the NExTLi workflow from imaging-guided region selection to molecularly resolved lipid profiles.
What NExTLi enables
- Structural and isomer-resolved lipidomics
- Low-input and single-cell/small-cell-population workflows
- Spatially informed lipidomics (imaging/MSI-guided)
- Tracer-based lipid fluxomics
- FAIR data delivery via the NExTLi Data & Analysis Portal
Why NExTLi
Lipid biology is central to health and disease but remains underexplored at a mechanistic level. Existing facilities generally support lipid profiling, but not the integrated structural characterization, spatial anchoring, and metabolic tracing required for mechanistic insight. A national landscape survey among 160 lipid researchers across Dutch academic centers confirmed strong demand (93%) for a dedicated national lipidomics infrastructure. NExTLi addresses this gap by combining these capabilities within a single coordinated workflow, with standardized QC and expert-supported interpretation.
How it works
- Intake: research question, samples, constraints, desired outputs
- Expert-guided experimental design and feasibility assessment
- Standardized sample processing, measurement, and QC
- Expert-supported data analysis and biological interpretation
- FAIR-by-design data packaging via the NExTLi Data & Analysis Portal
Three capability nodes
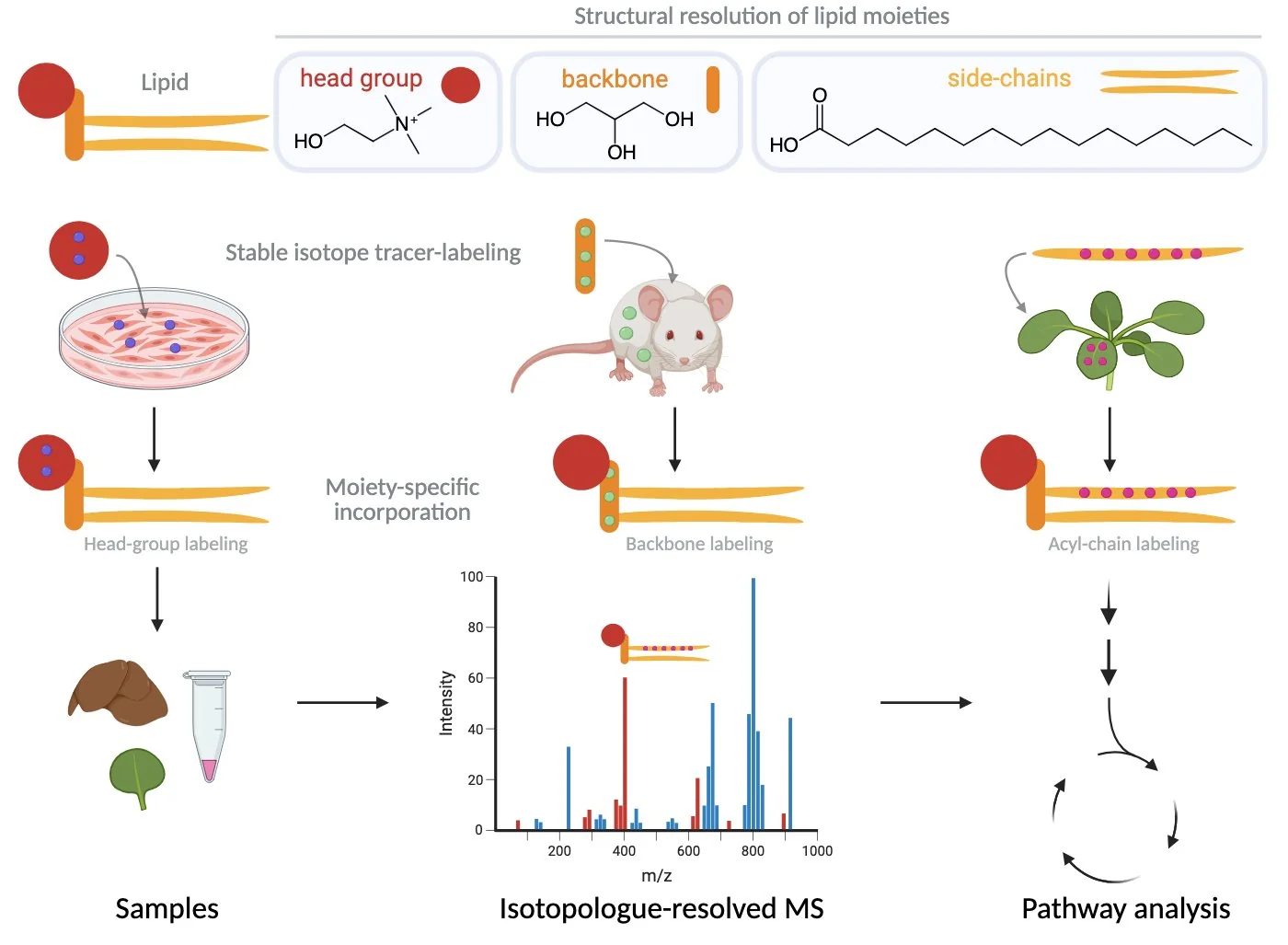
NExTLi is organized into three coordinated capability nodes that together cover the full analytical chain — from spatially defined sample preparation through in-depth molecular analysis to FAIR data integration. The tracer-based lipidomics workflow, illustrated below the node overview, shows how stable isotope labeling connects biological systems to pathway-level lipid dynamics.
Sample workflow & preparation
Spatially informed sampling, imaging-guided region selection, laser microdissection, and standardized low-input sample preparation. Connecting tissue architecture and cellular context to downstream molecular analysis.
Advanced lipidomics & fluxomics
High-resolution structural lipidomics, isomer resolution, and tracer-based metabolic flux analysis using next-generation nano-LC–MS/MS platforms. Molecular specificity from bulk samples to single-cell-scale material.
Data integration & analysis
FAIR-by-design data management, lipid-specific annotation and interpretation, statistical analysis, and the NExTLi Data & Analysis Portal — supporting reproducible workflows and interoperability with national data infrastructures.
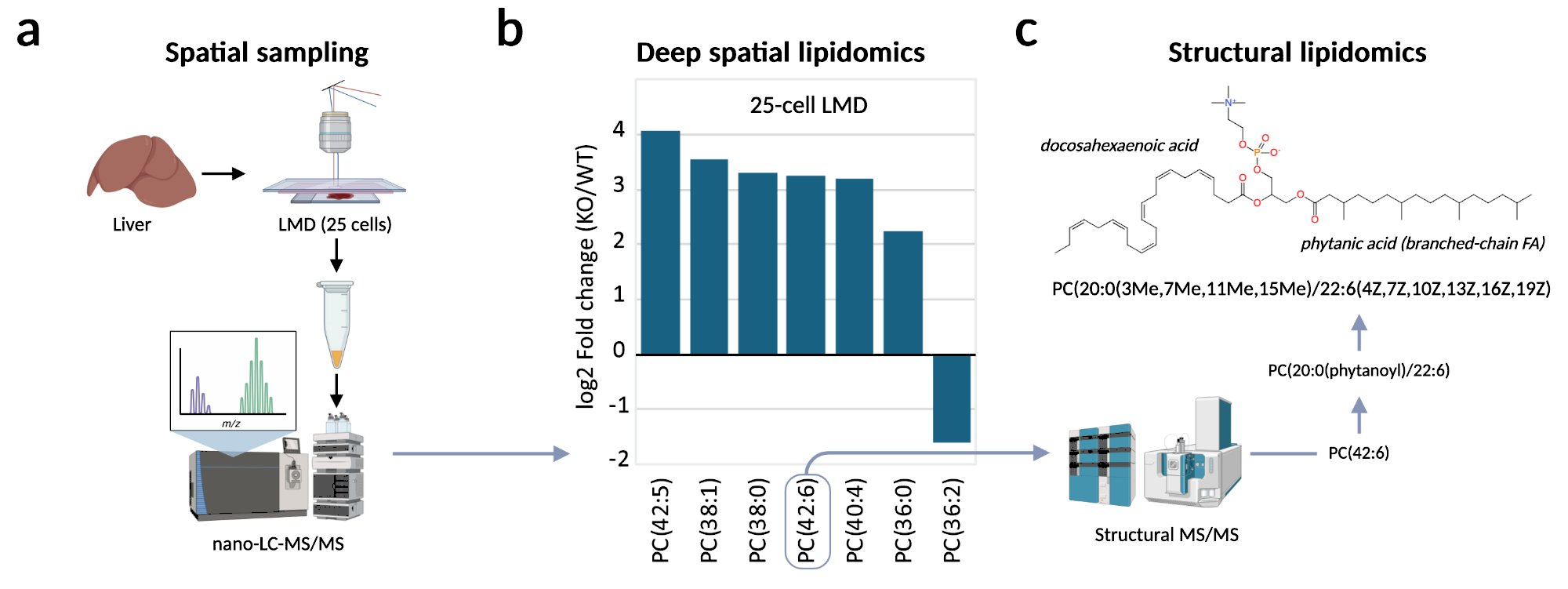
The figure below shows a proof-of-principle demonstration of the NExTLi spatial and structural lipidomics workflow. Liver tissue from a genetically defined metabolic disease model was sampled at ~25-cell resolution using laser microdissection, followed by nano-LC–MS/MS analysis. The resulting phosphatidylcholine profile (b) reproduces the same lipid remodeling signature previously observed in bulk liver homogenates — demonstrating that defined lipid phenotypes can be resolved at near single-cell scale. High-resolution MS/MS fragmentation (c) then resolves the specific molecular species driving that signature, identifying incorporation of the branched-chain fatty acid phytanic acid into a phosphatidylcholine species at full structural annotation level.
NExTLi is designed for interoperability with national research infrastructures including NL-BioImaging, X-omics, and Health-RI, and aligns with international standards through the Lipidomics Standards Initiative and ELIXIR.