Data & FAIR
NExTLi focuses on interpretable lipid biology. That requires data that stays usable: with QC evidence, annotation confidence, and metadata that survives beyond a single paper.
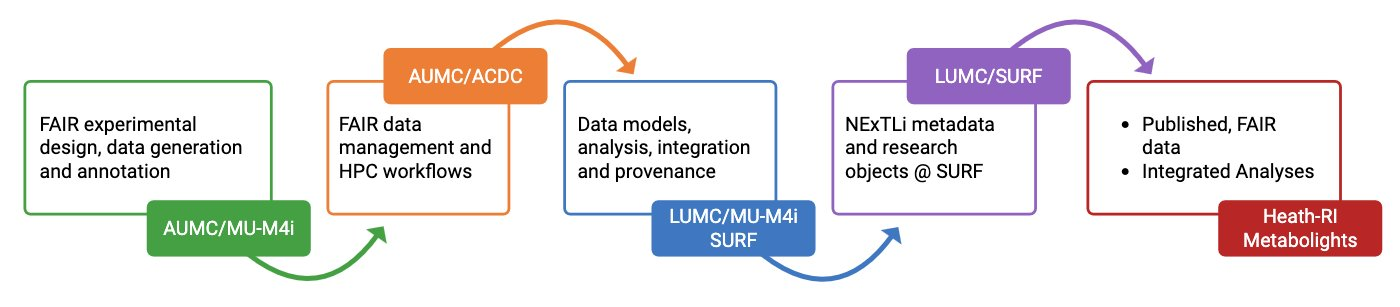
FAIR is not a checkbox at the end. The NExTLi Data & Analysis Portal — built on iSODA architectural principles and supported by institutional compute and storage infrastructure (ACDC, SURF) — embeds FAIR compliance into every step of the workflow, from experimental design to data delivery and long-term preservation. The diagram below shows how data flows through the NExTLi infrastructure from generation to published, reusable outputs.
What “FAIR” means here
- QC evidence and method context, not just files
- Annotation confidence metadata alongside results
- Structured metadata using ISA framework and community standards (Lipidomics Standards Initiative)
- Packaging designed for portal ingestion and repository deposition (e.g. MetaboLights)
Core deliverables
Deliverables are workflow-dependent, but the default is: results + QC + context + provenance. Users receive structured, reproducible research objects that can be explored interactively through the NExTLi Data & Analysis Portal or accessed locally.
- Analysis-ready tables with identifiers and annotation confidence metadata
- QC summaries (batch structure, controls, and key checks)
- Method and processing documentation with full provenance tracking
- Statistical results, visualization outputs, and structured research objects
What we capture early
Most “FAIR problems” are created before measurement. Mandatory metadata completion during project intake ensures experiments are FAIR-by-design from the start.
- Study design context (comparisons, covariates, constraints, intended outputs)
- Sample description and handling constraints (including low-input considerations)
- Workflow choice and QC plan (what success looks like)
- Reporting and reuse intent (integration targets, repository deposition, embargo needs)
What you can expect
Interpretability
Results come with context: QC evidence, annotation confidence, processing documentation, and expert-supported guidance on biological interpretation.
Reusability
Datasets are structured for reanalysis, cross-study comparison, and multi-omics integration. Formats conform to community standards suitable for deposition in established repositories.
NExTLi facilitates and encourages data sharing in accordance with FAIR principles: as open as possible, as closed as necessary. Decisions on data release, embargo periods, and access conditions remain with the data owners.