Infrastructure
Three coordinated capability nodes — sample workflow and preparation, advanced lipidomics and fluxomics, and data integration and analysis — integrated into a single end-to-end infrastructure for mechanistic lipid biology.
NExTLi is not a single instrument. It is an expert-supported infrastructure that combines standardized workflows, advanced mass spectrometry platforms, and FAIR data delivery so that results remain interpretable and reusable beyond one project. Spatial and non-spatial entry routes converge within a single coordinated workflow, as shown in the diagram below.
Analytical platforms
- High-sensitivity nano-LC–MS/MS for in-depth and low-input lipidomics
- Advanced nano-LC–MS/MS for structural characterization and isomer resolution
- Imaging-guided spatial sampling and laser microdissection
- NExTLi Data & Analysis Portal with FAIR-by-design workflows
- Complemented by existing Orbitrap, timsTOF, and MSI platforms
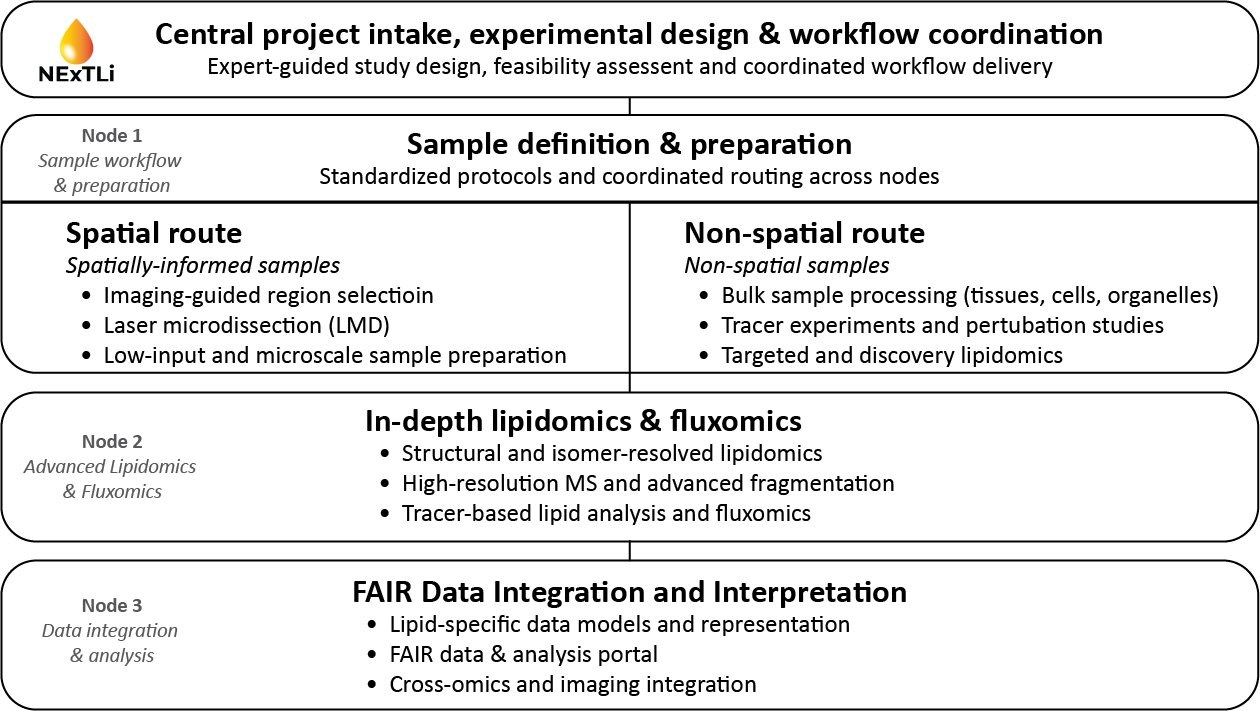
End-to-end workflow
NExTLi operates as a unified workflow from experimental conception and sample definition to structurally resolved lipid analysis and interpretable, reusable datasets. Centralized coordination ensures that distributed capabilities function as a single infrastructure from the user perspective.
- Central project intake, expert-guided study design, and feasibility assessment
- Coordinated routing across capability nodes
- Standardized sample handling, processing, QC, and documentation
- FAIR-by-design data delivery with annotation confidence metadata
What is standardized
Standardization ensures comparability and interpretability across projects. It covers the parts that most often break across labs and platforms, aligned with the Lipidomics Standards Initiative and community reporting standards.
- Sample handling guidance and documented constraints
- Processing steps, internal standards, and QC evidence
- Annotation confidence metadata and reporting templates
- FAIR packaging aligned to portal ingestion and community repositories
Three capability nodes
Sample workflow & preparation
Spatially informed sampling, imaging-guided region selection, laser microdissection, automated tissue processing, and standardized low-input sample preparation. Spatial and non-spatial entry routes converge within the infrastructure, ensuring that lipid measurements can be linked to defined biological structures.
Advanced lipidomics & fluxomics
High-resolution structural lipidomics and isomer resolution using next-generation nano-LC–MS/MS platforms, complemented by existing Orbitrap and ion-mobility-enabled systems. Tracer-based lipid fluxomics enables quantitative analysis of lipid synthesis, remodeling, and turnover. Low-input workflows extend these capabilities to single cells, small cell populations, and laser-captured tissue regions.
Data integration & analysis
The NExTLi Data & Analysis Portal provides standardized processing, visualization, annotation, statistical analysis, and FAIR data management. Building on existing frameworks (ACDC, iSODA, NeurolipidAtlas), this node ensures reproducible workflows, scalable compute, and interoperability with national data infrastructures.
NExTLi builds on established analytical platforms (Orbitrap Exploris 240, timsTOF Pro 2, spatial MS imaging systems) and will be extended with dedicated high-sensitivity and structural nano-LC–MS/MS instrumentation optimized for the workflows described above.